CUBE ChatShaala — Discussion Summary
CUBE ChatShaala — Discussion Summary
Date: 01 May 2026
Today’s CUBE ChatShaala session brought together two distinct but complementary threads of scientific inquiry. The first centred on Manali’s homelab experiment exploring the mutualistic relationship between Rhizobium bacteria and leguminous plants, set up on 30 April 2026. The second drew attention to a freshly published review paper — “Assessment of Venom Toxicity Using Drosophila melanogaster as a Model Organism” by Akanksha V. Joshi, Dr. Mrunal Ghag Sawant, and Sanika N. Parkar from the Haffkine Institute for Training, Research & Testing, Mumbai — published on 01 May 2026, the same day as this ChatShaala. Together, these two threads offered participants a rich experience of science at both the homelab and the published research levels.
published paper.pdf (242.6 KB)
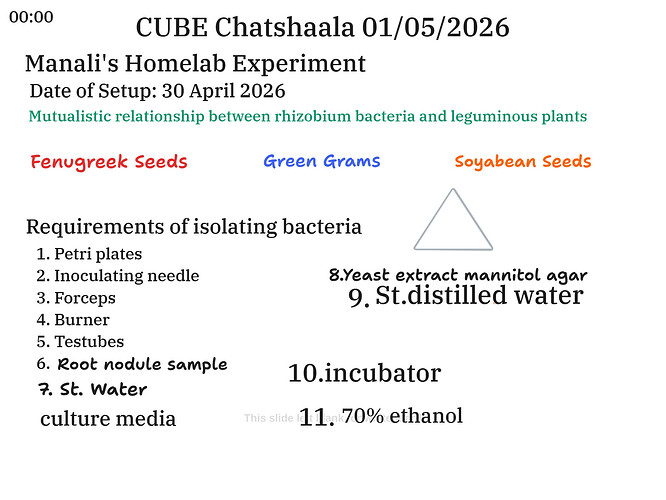
Part 1: Manali’s Homelab Experiment — Rhizobium and Leguminous Plants
The core scientific theme of this segment was the mutualistic relationship between Rhizobium bacteria and leguminous plants — one of the most significant biological partnerships known to agriculture and ecology. Manali chose three leguminous seeds for her experiment: Fenugreek seeds, Green Grams (Moong), and Soyabean seeds. Each of these plants forms root nodules in close association with Rhizobium species, making them ideal candidates for a home lab investigation of nitrogen fixation.
The group collaboratively built up a list of eleven essential requirements for isolating bacteria from root nodules: Petri plates, an inoculating needle, forceps, a burner, test tubes, a root nodule sample, sterilised water, a culture medium, Yeast Extract Mannitol Agar (YEMA), sterilised distilled water, an incubator, and 70% ethanol. Each item on this list was discussed in relation to its specific function — from aseptic technique (burner, ethanol) to supporting the slow growth of Rhizobium (YEMA medium, incubator).
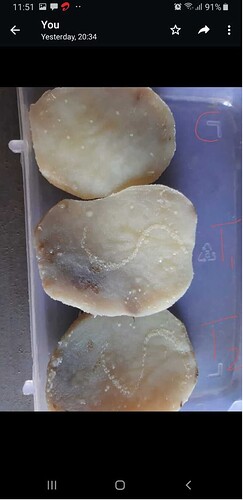
The second shared image — three boiled potato slices in a labelled plastic container showing varying degrees of microbial growth — connected the session directly to CUBE’s well-established tradition of homelab microbiology using alternative media. This method, pioneered within the CUBE network by Enas Fatima and collaborators, including Batul Pipewala, Aditya Joshi, and others, demonstrates that bacteria can be isolated and grown on boiled potato slices using the same streaking principles as formal laboratory agar plates. The potato provides starch and moisture as nutrients, and colonies that develop along the inoculation streak are distinguishable from environmental contaminants on the uninoculated control slice.
The group discussed how the potato medium relates to YEMA — whether it could serve as an accessible alternative for Rhizobium work, what controls are necessary, and how colony morphology can be described and documented even without advanced instrumentation.
Part 2: Published Review — Venom Toxicity and Drosophila melanogaster
The paper reviewed the toxicological effects of venoms from six major groups — snakes, honey bees, marine organisms, wasps, scorpions, and spiders — all studied using Drosophila melanogaster as the model organism.
The authors opened by establishing why Drosophila is such a powerful model: approximately 60% of its genome is homologous with the human genome, and nearly 75% of human disease-associated genes have functional counterparts in the fly. Its short life cycle, ease of culture, and well-characterised genetics make it ideal for toxicological and neurological research.
Key findings discussed across the venom categories included:
Snake venom (Naja naja): Rather than showing only toxic effects, Naja naja venom demonstrated some paradoxically beneficial outcomes in fly models — improving cognitive behaviour, reducing reactive oxygen species (ROS), and counteracting the oxidative damage caused by the carcinogen NDEA. Venom components showed anti-proliferative and anti-angiogenic properties, suggesting potential anti-cancer applications.
Honey bee venom (Apis mellifera): Bee venom reduced adipokinetic hormone (AKH) levels in the fly’s central nervous system, disrupted antioxidant enzyme activity (SOD and CAT), and caused structural damage to muscle tissue. The venom peptide apamin, when expressed in Drosophila, unexpectedly showed antimicrobial activity localised in the gut, along with effects on intestinal stem cell proliferation. The peptide Minimine, at sub-lethal doses, caused larvae to survive but stop growing, producing adults one-fourth normal size, whose offspring were entirely normal.
Marine venoms (Physalia physalis and terebrid snails): Marine venoms altered movement, circadian rhythm, and heat-avoidance behaviour. Two peptides from terebrid marine snails had opposing effects — one reduced the fly’s response to heat-induced pain (antinociceptive), while the other increased food intake. These findings support the use of Drosophila behavioural assays as a screening platform for novel bioactive marine compounds.
Wasp venoms: Parasitic wasps such as Leptopilina boulardi, L. Heterotoma, and Pachycrepoideus vindemiae use venom to suppress the fruit fly’s immune system, allowing wasp larvae to develop inside the host. Venom proteins called venosomes enter fly immune cells through lipid raft-dependent pathways and deliver disruptive cargo. The wasp Asobara japonica was found to degrade imaginal discs — the larval precursor tissues for adult organs — causing delayed development. The venom of P. Vindemiae was shown to kill fly immune cells through programmed cell death (apoptosis).
Scorpion venoms: Drosophila was found to be generally less sensitive to scorpion venoms than other insect models, but notable effects were still observed. The Hottentotta judaicus venom reduced locomotion and increased sleep time in flies at sub-lethal doses. The C56 toxin from Mesobuthus tamulus (the Indian red scorpion) enhanced nerve–muscle communication at the larval neuromuscular junction by promoting calcium ion entry through NMDA-type receptors.
Spider venoms: Multiple spider venom peptides were studied, with effects ranging from presynaptic calcium channel blockade (spider Hololena curta) to massive neurotransmitter release stimulation (α-latrotoxin from the black widow, Latrodectus mactans). The concept of the Toxic Male Technique (TMT) — where genetically engineered males deliver venom proteins to females during mating, reducing female lifespan — was introduced as a novel, venom-based pest control strategy.
The paper concluded that Drosophila melanogaster is a reliable, accessible, and ethically appropriate model for investigating the biological activity of diverse venoms, with implications for medicine, neuroscience, and pest management.
 Provocative Questions
Provocative Questions
From Manali’s Homelab Experiment:
-
Fenugreek, green grams, and soyabean all harbour Rhizobium, but they associate with different strains. If nodule samples from all three plants were plated on the same YEMA plate, would the resulting colonies be morphologically distinguishable from one another? How would you design an experiment to test this?
-
The whiteboard lists “St. Water” (item 7) and “St. distilled water” (item 9) as separate requirements. Are these genuinely distinct in function, or is this an oversight in the protocol design? What specific steps in root nodule isolation require each type of water, and why does the grade of water matter?
-
The boiled potato slice method has been validated for isolating bacteria from curd, milk, and soil in CUBE homelabs. Could it support the growth of slow-growing, fastidious bacteria like Rhizobium, which normally requires the specialised YEMA medium? What evidence would you look for to confirm or refute this?
-
If a control potato slice (uninoculated) shows scattered microbial growth while the inoculated slice shows growth along the streak, what does this tell us about the experiment? At what point does the level of contamination on the control make the entire experiment uninterpretable?
-
How would you confirm that the colonies you have isolated from a root nodule are genuinely Rhizobium and not other root-associated endophytes? What is the most accessible confirmation test available in a home lab setting?
From the Venom Toxicity Review Paper:
-
Naja naja venom showed cognitively beneficial effects and reduced oxidative stress in fly models, yet it is simultaneously one of the most lethal venoms known to humans. What does this apparent paradox tell us about the dose-dependency and context-specificity of venom action? Does a beneficial effect in a fly model translate meaningfully to a therapeutic signal for human disease?
-
The venom peptide Minimine produced larvae that survived but stopped growing, becoming one-fourth the normal adult size — yet their offspring were entirely normal. Does this mean Minimine’s effect was purely physiological rather than genetic? What biological mechanism might explain a sub-lethal venom effect that does not pass to the next generation?
-
Parasitic wasp venoms have evolved specifically to suppress Drosophila’s immune system — essentially hijacking the fly’s biology from the inside. What does this tell us about the co-evolutionary arms race between host immunity and parasite venom? And could studying these venom proteins reveal new targets for human immunosuppressive therapies?
-
The Toxic Male Technique uses venom proteins transferred during mating to reduce female lifespan by 37–64%. What are the ecological risks of releasing such genetically engineered insects into the wild? Could there be unintended consequences for non-target species?
-
Drosophila is described as “less sensitive” to scorpion venoms than other insect models. What does sensitivity mean in this context — is it about receptor compatibility, metabolism, body size, or something else? And does lower sensitivity make Drosophila a less useful model for scorpion venom research, or does it open different kinds of experimental questions?
 What I Have Learned
What I Have Learned
This session layered two very different scales of scientific work on top of each other, and that layering was genuinely instructive.
From Manali’s homelab work, I was reminded that the most important thing in any experiment is not the sophistication of the equipment but the clarity of the biological question. The question here — can we isolate the bacterium responsible for nitrogen fixation from the root nodules of leguminous plants using accessible materials? — is both scientifically meaningful and practically achievable. The eleven-item requirements list is not a formality; it represents a coherent experimental logic in which every tool has a defined role.
I also came away with a deeper appreciation for Yeast Extract Mannitol Agar as a selective tool. The medium does not just feed Rhizobium — it does so selectively, exploiting the organism’s specific metabolic preferences to give it an advantage over faster-growing contaminants. This principle of selective media is one of the foundational ideas in microbiology, and seeing it applied in a homelab context made it more tangible than any textbook explanation.
From the venom paper, the most significant conceptual takeaway for me is the idea of Drosophila as a universal biological translator. Because so much of its genome is conserved with mammals — and because its behaviours are observable, quantifiable, and responsive to interventions — it can be used to decode the biological effects of compounds as varied as cobra venom and sea snail peptides. The fact that approximately 75% of human disease genes have functional counterparts in the fly is not merely a statistic; it is the foundation for an entire field of translational research.
The Toxic Male Technique struck me as a genuinely novel intersection of venom science and pest management — a reminder that basic research into venom mechanisms can produce unexpected applications with real-world consequences.
And across both discussions, I was struck by how much science depends on controls and comparisons. Whether it is an uninoculated potato slice sitting beside an inoculated one, or a fly exposed to venom alongside an unexposed sibling, the control is not just a procedural requirement — it is the thing that makes the experiment speak for itself.
TINKE Moments (This I Never Knew Earlier)
TINKE 1: Naja naja venom can improve cognition in flies.
Most participants would intuitively associate snake venom with harm and death. The TINKE moment here is learning that at specific doses and in specific genetic contexts, Naja naja venom actually improved behavioural parameters in Drosophila and counteracted oxidative damage. This challenges the flat assumption that “venom = toxin = bad,” and opens the door to thinking about venoms as libraries of bioactive molecules with dose-dependent, context-specific effects.
TINKE 2: Wasp venoms travel inside cellular vehicles called venosomes.
The idea that venom is simply injected as a liquid is intuitive but incomplete. The TINKE here is learning that parasitic wasp venoms can be packaged inside membrane-bound vesicles (venosomes), which then enter the fly’s immune cells through a specialised lipid raft-dependent endocytic pathway. This is not crude poisoning — it is a molecularly sophisticated delivery system evolved over millions of years.
TINKE 3: A sub-lethal dose of bee venom peptide can produce dwarf flies whose offspring are normal.
This is a TINKE that challenges naive assumptions about inheritance and development. Minimizing at the LD50 dose produced flies one-fourth the normal size — yet those flies’ offspring were entirely normal. This implies the venom acted on developmental physiology without altering the germline. Understanding this distinction between somatic and germline effects is a genuinely important conceptual boundary.
TINKE 4: YEMA is not a generic medium — it is ecologically specific to Rhizobium.
For participants unfamiliar with selective media, listing “Yeast Extract Mannitol Agar” might seem like picking any convenient agar. The TINKE is realising that YEMA is specifically formulated to support Rhizobium’s slow, nitrogen-fixing metabolism — and that using a general nutrient agar could result in the target organism being completely outcompeted by faster-growing contaminants, making the experiment fail silently.
TINKE 5: Boiled potato slices are a validated scientific platform, not an informal substitute.
Participants encountering CUBE’s homelab microbiology tradition for the first time often dismiss potato slice media as improvised. The TINKE is learning that this method has a documented track record of successfully isolating bacteria from milk, curd, and other sources — with proper controls, described colony morphology, and reproducible results reported across multiple CUBE homelabs. Accessibility and rigour are not mutually exclusive.
 Gaps and Misconceptions
Gaps and Misconceptions
Gap 1: Temporal feasibility of the experiment is not addressed.
Manali’s setup date is 30 April 2026 — one day before this session. Root nodule formation in leguminous plants typically requires the plant to germinate, develop a root system, and then undergo the infection and nodulation process, which takes a minimum of two to three weeks. The session did not clarify whether Manali already had nodulated plants from a prior setup, or whether the 30 April date marks the beginning of a longer, multi-week experiment. This ambiguity could mislead newer participants into thinking bacterial isolation begins within days of sowing.
Gap 2: Colony verification methods are absent from the experimental plan.
The whiteboard covers the setup and culture phase thoroughly, but no method for confirming that isolated colonies are actually Rhizobium is mentioned. Gram staining, motility testing, and especially a re-inoculation nodulation test — where isolated bacteria are applied to seedlings and checked for nodule formation — are the standard verification steps. Their absence from the listed requirements is a notable gap.
Gap 3: The venom paper does not address interspecific variation within Drosophila.
Almost all studies in the paper use Drosophila melanogaster, yet one study on spider venom used Drosophila suzukii, a pest species with different ecological characteristics. The paper does not critically examine whether findings from D. melanogaster are generalisable across Drosophila species, which is a limitation worth flagging for participants who might assume all fruit flies respond identically.
Gap 4: The potato slice image lacks clear experimental labelling.
The second image shows three slices with partial red annotations, but no legend or key was provided during the session. It is impossible to determine from the image alone whether the three slices represent different seed sources, different inoculum concentrations, or independent replicates. This gap in documentation weakens the image as experimental evidence.
Misconception 1: All legumes associate with the same Rhizobium species.
The three plants chosen by Manali — fenugreek, green grams, and soyabean — do not all associate with the same bacterial partner. Soyabean, for instance, associates primarily with Bradyrhizobium japonicum, while the other two associate with distinct Rhizobium strains. Treating these as equivalent sources of the same organism could produce confusing results when colony morphologies differ across plant species.
Misconception 2: A beneficial venom effect in Drosophila implies a safe or therapeutic effect in humans.
The discussion of Naja naja venom improving cognition in fly models could inadvertently create the impression that snake venom has straightforward medical applications. In reality, the translation from Drosophila models to human therapeutics involves enormous biological distance and requires rigorous dose-response characterisation, toxicity profiling, and clinical validation. The fly model is a starting point for hypothesis generation, not a proof of human efficacy.
Misconception 3: Venom toxicity is a simple, linear dose–response relationship.
Several studies in the paper showed non-linear or unexpected effects — dwarf flies surviving sub-lethal doses, venom improving rather than harming behaviour, and immune suppression rather than direct lethality in wasp venom work. This complexity should caution participants against assuming that “more venom = more harm” in all contexts.